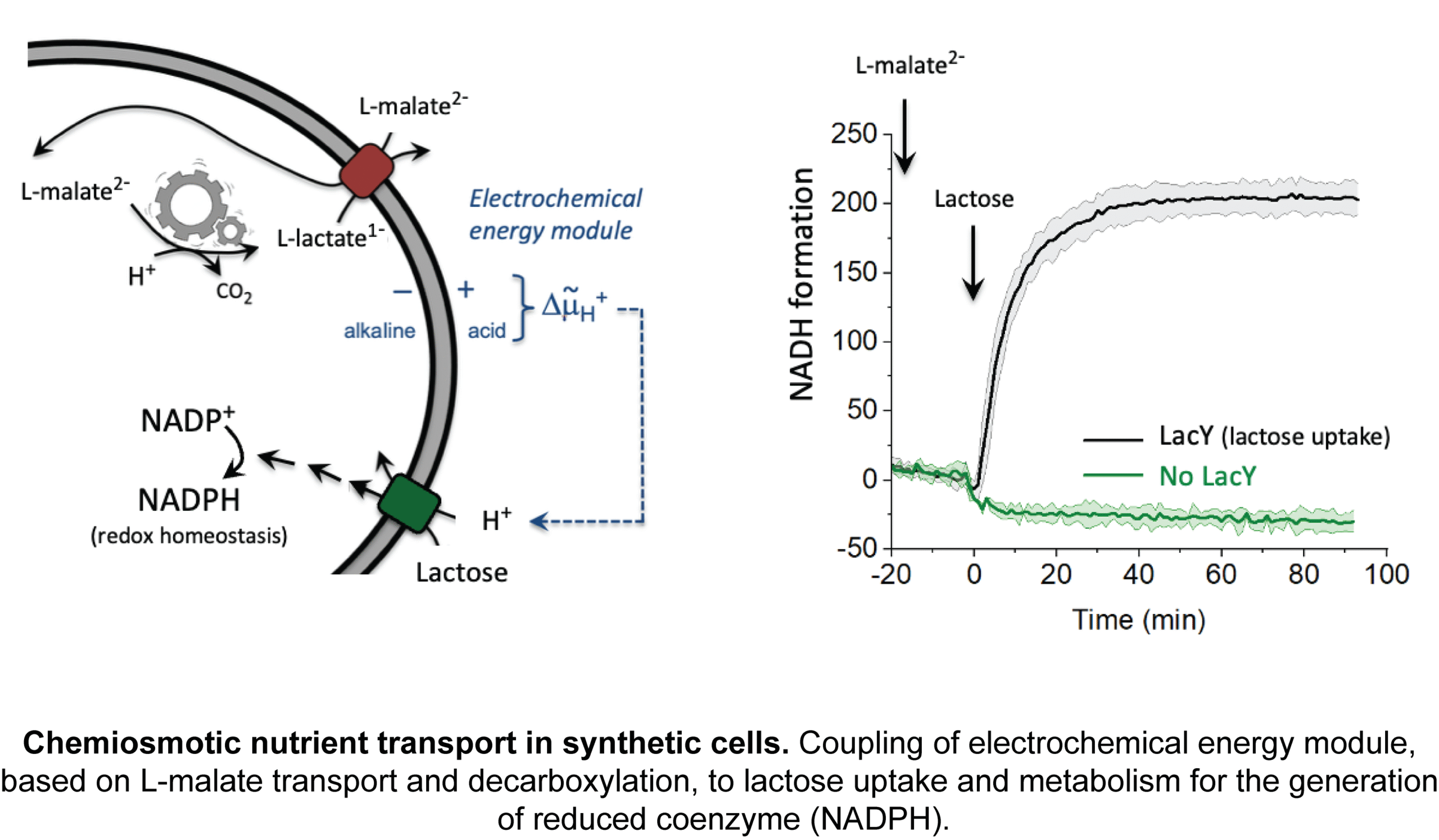
Cell volume regulation and physicochemical homeostasis
Bacteria maintain an outward osmotic pressure, or turgor, to grow and divide. Biological membranes are semipermeable, and changes in the external osmolality cause transmembrane water fluxes that either lead to loss of turgor, dehydration and ultimately plasmolysis, or increased turgor and possibly cell lysis. Managing cell volume homeostasis is particularly important for (micro)organisms that are exposed to osmotic fluctuations. We study the osmoregulatory ABC transporter OpuA and the membrane-bound cyclase for 2nd messenger cyclic di-AMP CdaA, which are involved in cell volume regulation and physicochemical homeostasis.
Relevant publications: doi: 10.1038/s42003-024-07420-x; doi: 10.1128/mmbr.00181-23; doi: 10.1126/sciadv.abd7697; doi: 10.1021/acs.chemrev.3c00622
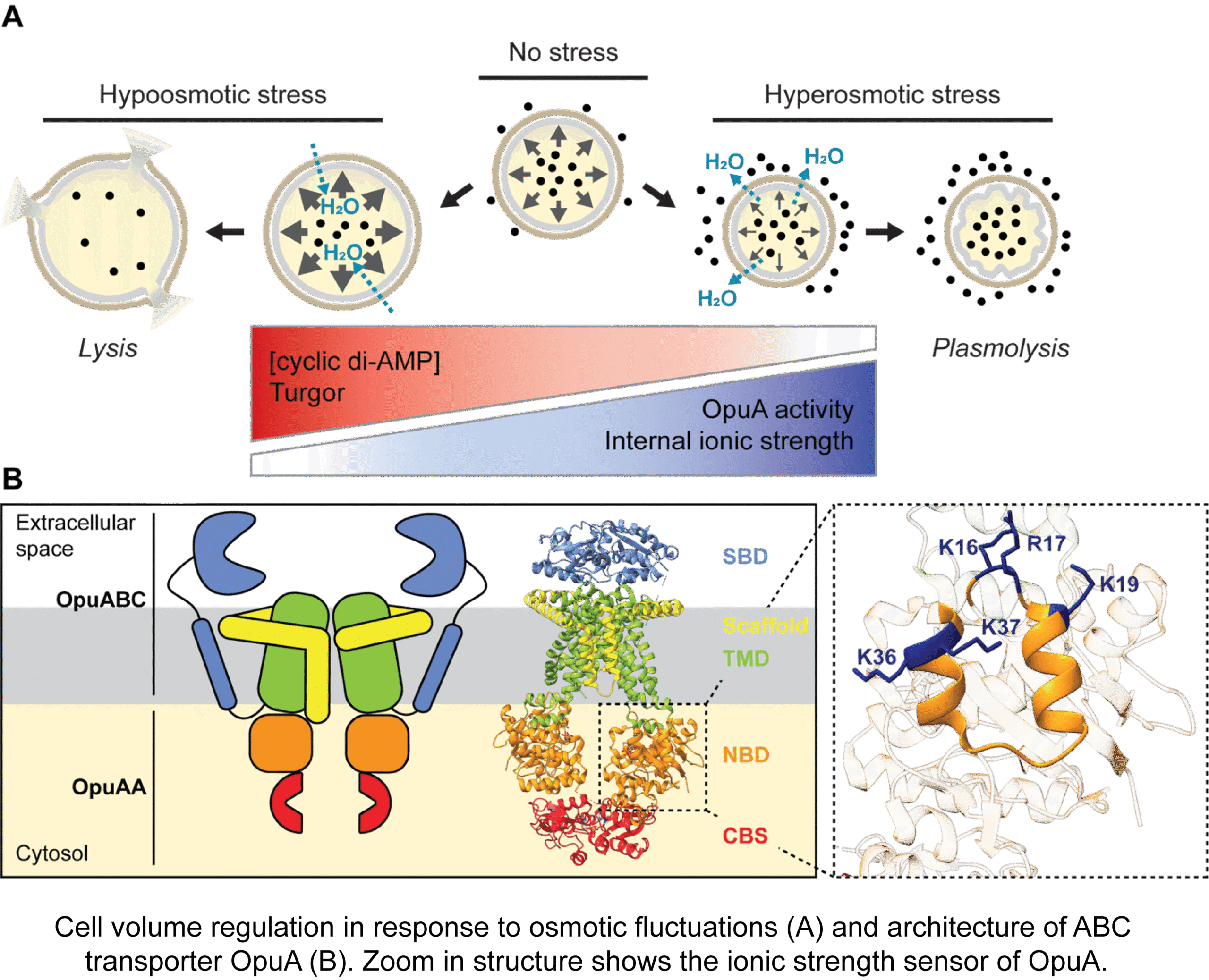
Dynamic Organization and Protein Diffusion in Bacterial Cells
Spatial heterogeneity in composition and dynamics of (macro)molecules, both in the cytoplasm and in cellular membranes, forms the basis of functional interactions and reactions in the cell. State-of-art optical microscopy techniques [including particle and single-molecule tracking, displacement mapping, fluorescence recovery after bleaching bleaching (FRAP), and photoactivated localization microscopy (PALM)] are used in combination with functional characterization and theoretical modeling to elucidate the chemistry and physics of the cell. We study how environmental factors influence protein mobility and localization in the cytoplasm, periplasm and inner membrane of bacterial cells. By linking molecular properties to cell-scale architecture, this work reveals how non-uniform intracellular environments regulate biochemical processes in living cells. The resulting insights are important not only for fundamental understanding of cellular organization but also for the design of cell-mimicking systems with desired properties.
We are also looking for PhD students with a background in (bio)physics, biochemistry or and (microbial) cell biology to participate in this challenging research.
Relevant publications: 10.1016/j.mib.2025.102635, 10.1126/sciadv.abo5387, 10.1038/s42003-024-06216-3, 10.1038/s41467-026-69951-7, 10.1016/j.jmb.2023.168420
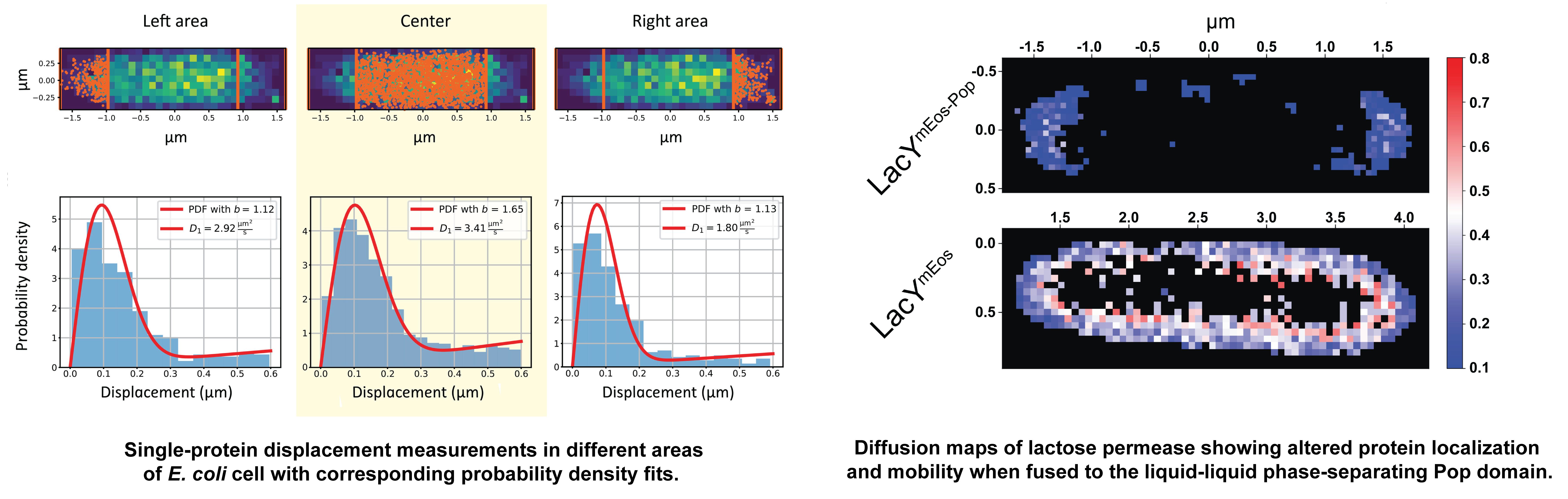
Evolution and engineering of membrane transport
We study plasma membrane transporters of yeast, and implement state-of-the-art optical microscopy (super-resolution optical microscopy, single-molecule tracking) to determine the traffic and localization of newly synthesized proteins. We have shown that the substrate spectrum of five wild-type yeast amino acid transporters is substantially broader than previously described, and by applying evolutionary pressure, single mutations emerged in the population and transporters with a gain of function were obtained.
Relevant publications: doi: 10.7554/eLife.93971; doi: 10.1038/s41467-018-02864-2.
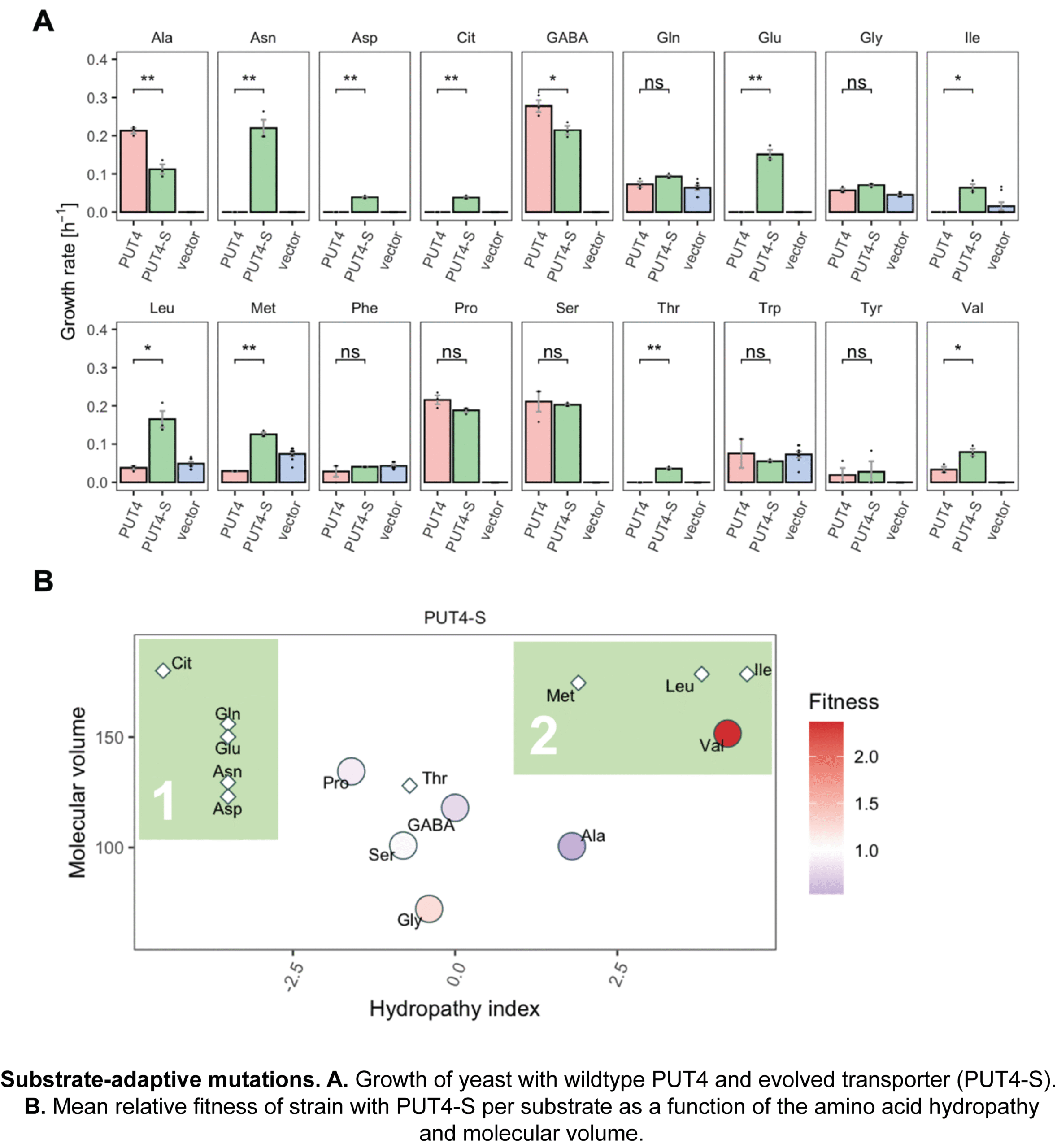
Bottom-up construction of synthetic cells
Living systems are enormously complex, and even the engineering of the smallest living systems is challenging as cells are still largely a black box. Non-linear interactions between metabolic and regulatory networks, system redundancy, emergent properties, unknown mechanisms, evolutionary pressures, context-specific behavior and other factors resist easy manipulation. We build artificial life-like systems from known components and study their emergent properties. The ultimate goal is to develop robust scalable synthetic platforms to unlock new scientific insights and drive transformative innovations across health, biotech industry and the environment.
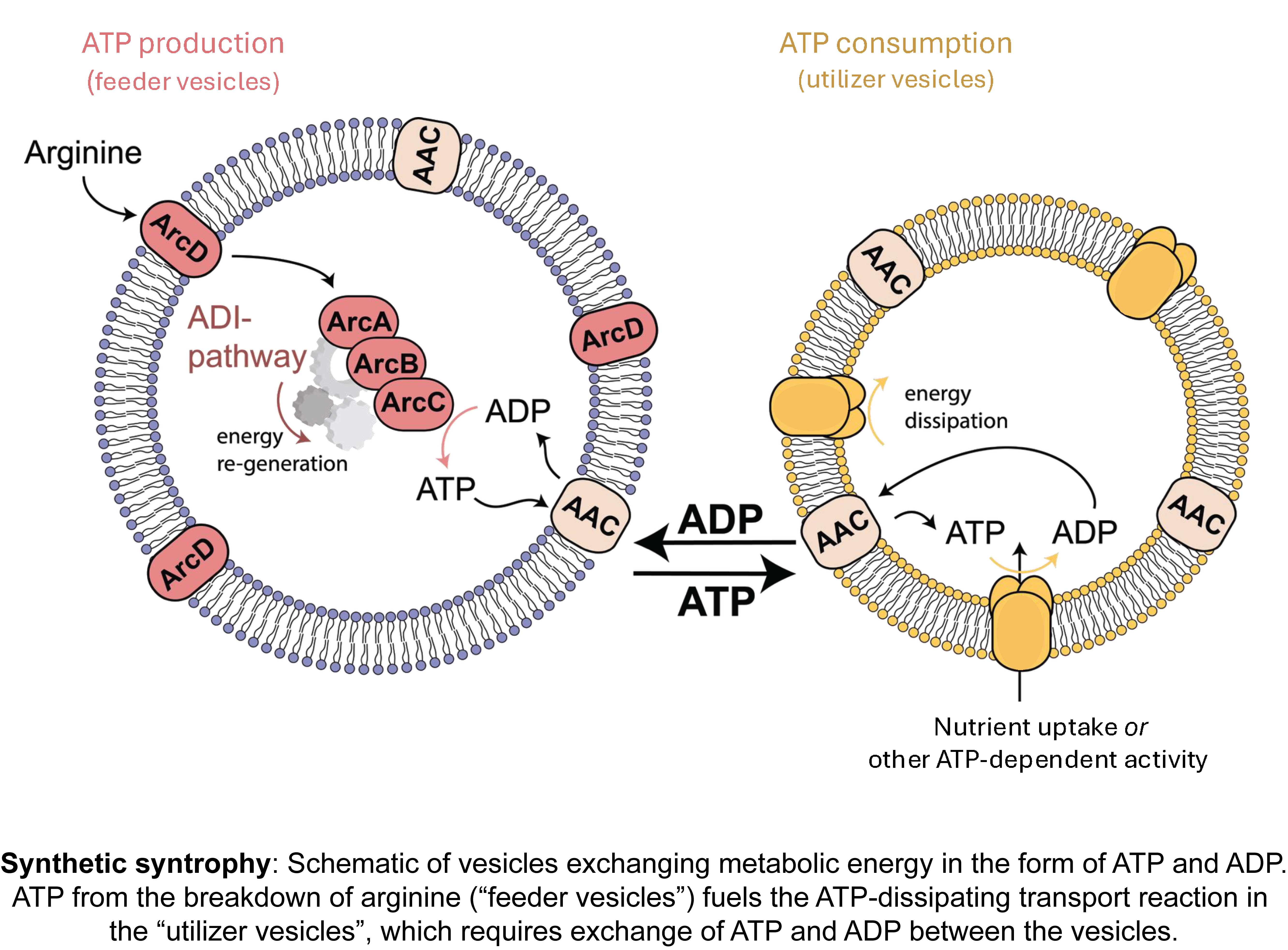
We focus on creating synthetic cells that can sustain themselves, grow, and ultimately divide. This requires, at a minimum, compartmentalization of reaction networks, conversion of nutrients into metabolic energy, an information processing system, and a division mechanism. We combine experimental and computational approaches to create controllable, life-like systems. Recent achievements are the construction of synthetic organelles that exchange metabolic energy (synthetic syntrophy) and electrochemical modules to drive solute transport and metabolism.
Relevant publications: doi: 10.1038/s41565-024-01811-1; doi: 10.1038/s41467-024-52085-z; doi: 10.3791/66627; doi: 10.1021/acssynbio.4c00073; doi: 10.1021/jacsau.1c00406; doi: 10.1038/s41596-022-00734-2
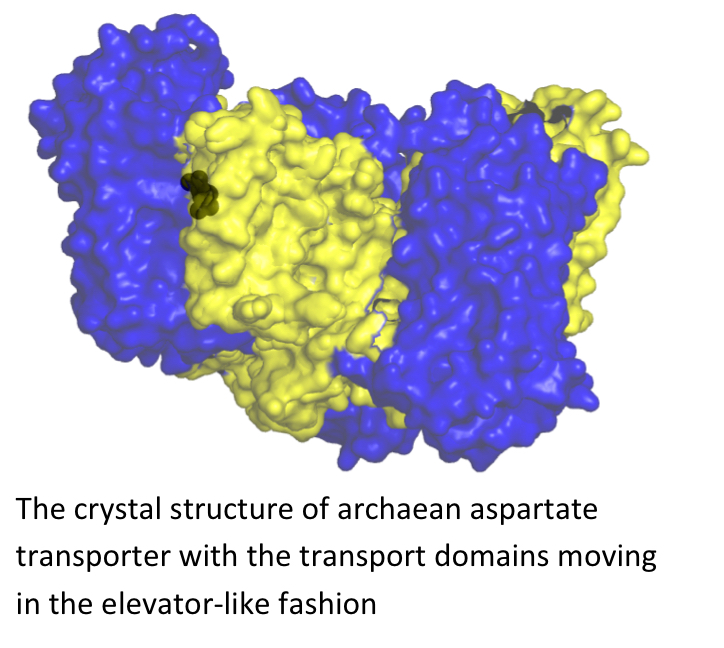
Aspartate (Glutamate) transport across the membrane
Glutamate transporters are secondary transporters found in many eukaryotes, bacteria and archaea. In prokaryotes these transporters mediate the uptake of nutrients, whereas in mammals, where glutamate is a major excitatory neurotransmitter, they catalyze the neurotransmitter reuptake into neurons and glial cells that follows synaptic signal transduction. Maintaining a low extracellular concentration of the neurotransmitter is essential for sensitive signal transmission and for the prevention of neurotoxicity. The transporters couple the uptake of glutamate to the symport of cations: three sodium ions and one proton. We use the archaean aspartate transporters as the model structures for mammalian glutamate transporters. We are interested both in the structural elucidation of different states of these transporters (by means of X-ray crystallography) and also in their dynamics, which we study with EPR and single molecule FRET.
The novel player in the team – ECF transporters in ABC superfamily
Energy-coupling factor (ECF) transporters belong to the ATP-binding cassette (ABC)-transporter family and mediate the uptake of essential micronutrients in many prokaryotic species. ECF transporters are present in approximately 50% of prokaryotic species, and that they catalyse the cellular uptake of a range of micronutrients, including water-soluble vitamins (such as riboflavin and thiamin) and their precursors, as well as transition metal ions. We are studying these transporters by combination of biochemical, biophysical and crystallography tools to unravel their transport mechanism.
Vitamin transport in bacteria
Pnuc transporters catalyze cellular uptake of the NAD+ precursor nicotinamide riboside (vitamin B3) and belong to a large superfamily that includes the Sweet sugar transporters. The latter use a facilitated-diffusion mechanism to catalyze low-affinity sugar transport and are particularly abundant in plants, in which they allow, for instance, nectar secretion. In humans, the SWEET transporter might represent the elusive glucose transporter of the basolateral membrane of enterocytes in the intestine. PnuCs also use a facilitated-diffusion mechanism but linked to metabolic trapping in the bacterial cytoplasm by the specific kinase NadR, which converts NR into nicotinamide mononucleotide (NMN) and NAD+. Recently we have solved the crystal structure of PnuC from Neisseria mucosa, which adopts a highly symmetrical fold with 3 + 1 + 3 membrane topology not previously observed in any protein. Currently we are also investigating other transporters responsible for the vitamin uptake in bacteria.